Normal Twin PET - Personalized Baseline for Whole-Body PET Analysis
PhD Project by Christian Hinge
Whole-body PET/CT imaging is widely used for disease diagnosis and treatment assessment, but quantitative analysis is challenged by large interpatient variability caused by physiological FDG uptake patterns. Factors such as age, body composition, scanner type, and uptake time can obscure disease-related signals and limit diagnostic precision.
Project BackgroundThis study presents a novel deep learning approach for generating personalized normal twin PET images based on the patient's own CT scans. The synthetic normal twin represents how the patient’s PET image would appear under healthy conditions and serves as an individualized reference baseline. By comparing PET images with their normal twins, the method enables confounder correction and robust anomaly detection. The approach for generating the normal twin is illustrated in Figure 1.
Project PotentialThe developed model generates patient-specific normal PET images and supports quantitative correction of physiological variability. Initial results demonstrate a significant reduction of SUV variance and improved detection of abnormal uptake patterns without requiring manually annotated training data.
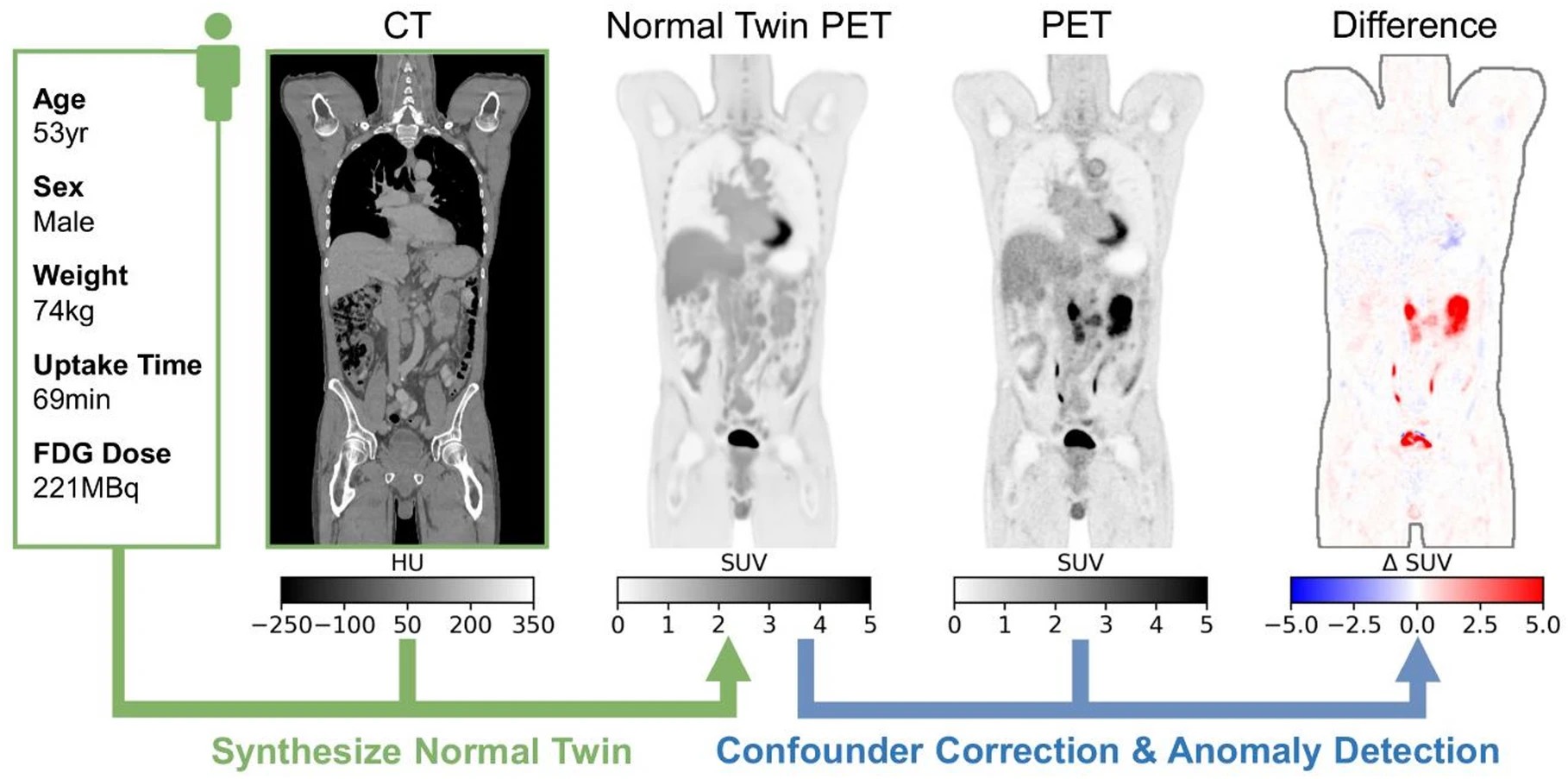
Figure 1: Normal Twin PET pipeline. Personalized normal PET images are generated from CT and patient attributes and used for confounder correction and anomaly detection.